What is XenonPy project
Note
To all those who have purchased the book Materials Informatics published by KYORITSU SHUPPAN:
The link to the exercises has changed to https://github.com/yoshida-lab/XenonPy/tree/master/mi_book. Please follow the new link to access all these exercises.
We apologize for the inconvenience.
Overview
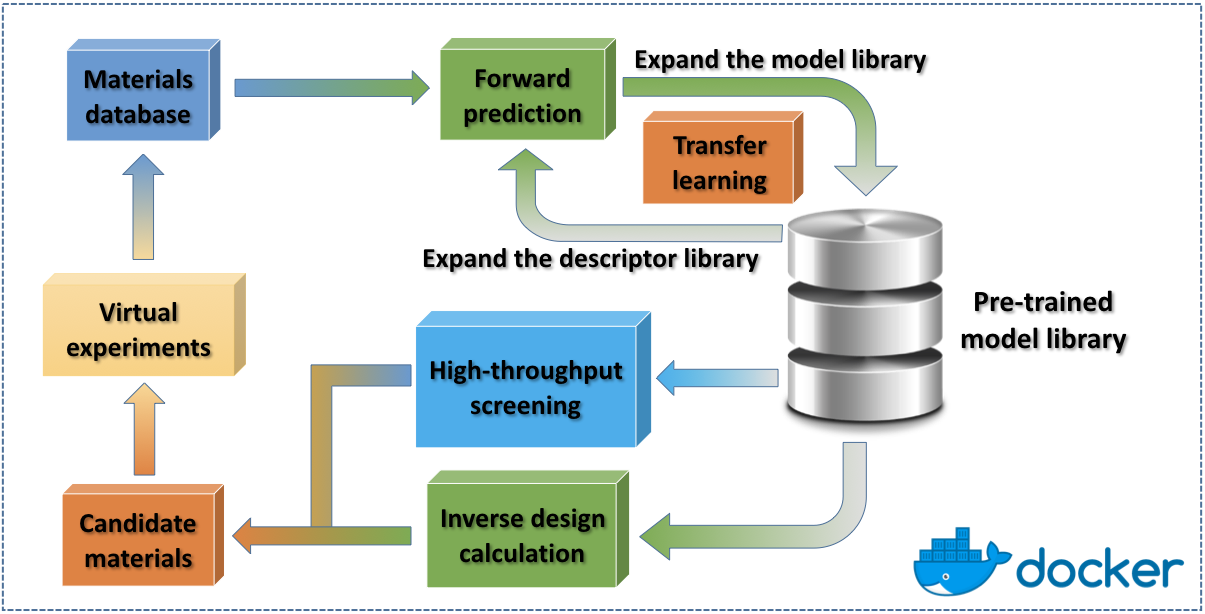
XenonPy is a Python library that implements a comprehensive set of machine learning tools for materials informatics. Its functionalities partially depend on Python (PyTorch) and R (MXNet). This package is still under development. The current released version provides some limited features:
Interface to the public materials database
Library of materials descriptors (compositional/structural descriptors)
pre-trained model library XenonPy.MDL (v0.1.0.beta, 2019/8/9: more than 140,000 models (include private models) in 35 properties of small molecules, polymers, and inorganic compounds) [Currently under major maintenance, expected to be recovered in v0.7]
Machine learning tools.
Transfer learning using the pre-trained models in XenonPy.MDL
Citation
XenonPy is an on-going research project that covers multiple important topics in materials informatics. We recommend users to cite the papers that are relevant to their specific use of XenonPy. Please refer to Features for details of each feature in XenonPy with its corresponding citation. User can also check the publication list below to pick the relevant citations.
Features
XenonPy has a rich set of tools for various materials informatics applications.
The descriptor generator class can calculate several types of numeric descriptors from compositional, structure.
By using XenonPy’s built-in visualization functions, the relationships between descriptors and target properties can be easily shown in a heatmap.
XenonPy also supports an interface to use the rdkit descriptors and provides the iQSPR algorithm for molecular design.
Transfer learning is an important tool for the efficient application of machine learning methods to materials informatics. To facilitate the widespread use of transfer learning, we have developed a comprehensive library of pre-trained models, called XenonPy.MDL. This library provides a simple API that allows users to fetch the models via an HTTP request. For the ease of using the pre-trained models, some useful functions are also provided.
See Features for details.
Sample
Sample codes of different features in XenonPy are available here: https://github.com/yoshida-lab/XenonPy/tree/master/samples
Publications
Contributing
XenonPy is an open source project inspired by matminer.
This project is under continuous development. We would appreciate any feedback from the users.
Code contributions are also very welcomed. See Contribution guidelines for more details.



